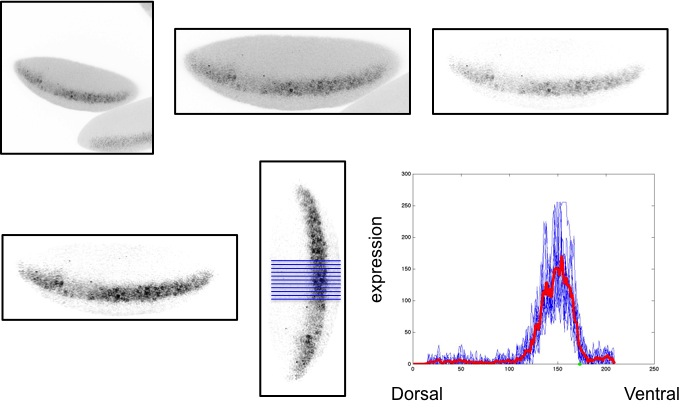
When fitting a mathematical model, regardless of the field in which the model is being applied, the quality of the data used is of tremendous importance. For models of transcriptional regulation, both spatial and temporal protein (TF) and mRNA concentrations need to be considered. To obtain the most accurate quantitative measurements of these concentrations, one needs to have a thorough understanding of where the data is coming from and what procedures can be implemented to assure that the data being analyzed is comparable on all levels.
Using raw data from microscope images in modeling gene expression levels presents the researcher with many challenges; one must be able to remove any extraneous information from the image and be able to compare images prepared and taken from different embryos on different days. For this reason, we focus much attention on processing these images, including noise and background subtraction, normalization, spatial registration, and extraction of quantitative levels of gene expression.