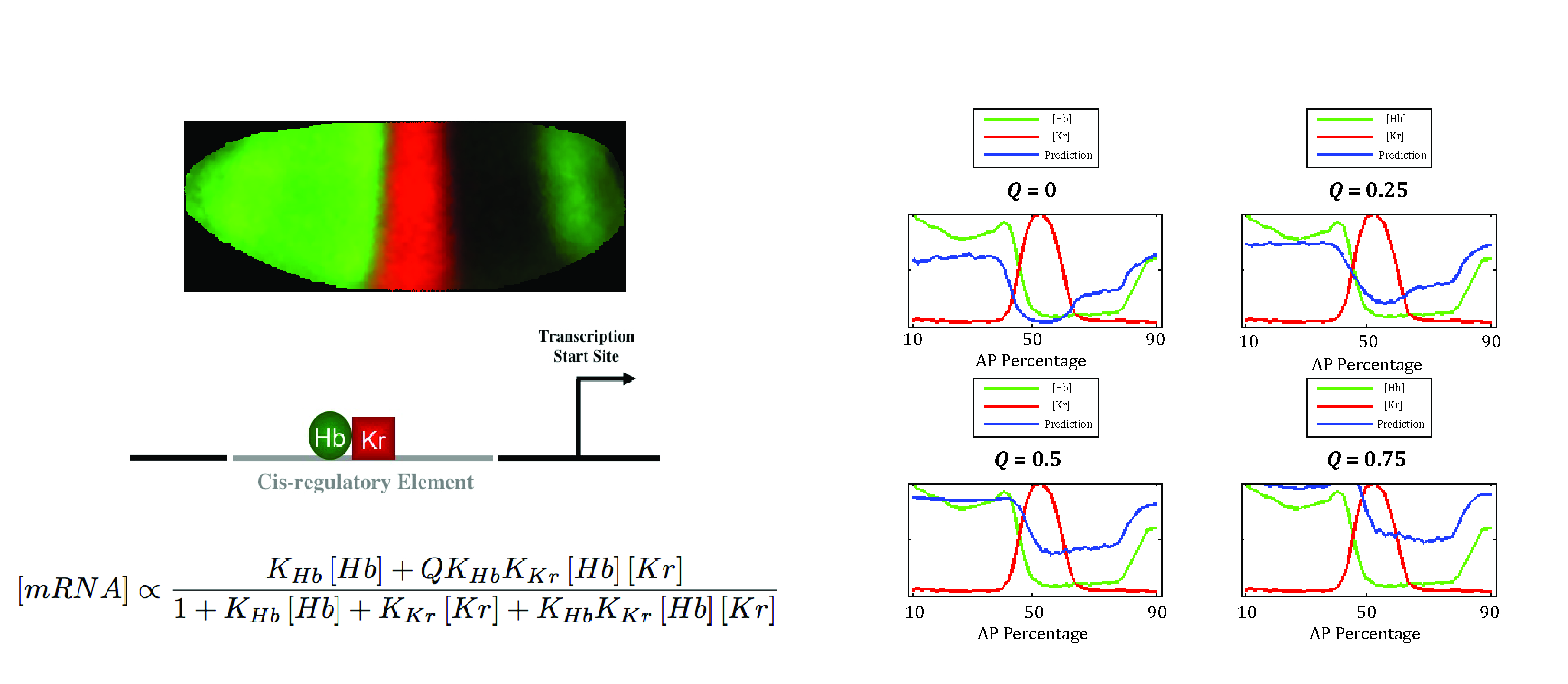
When implementing a mathematical modeling approach, working with endogenous enhancers can often be difficult due to the complexity of the enhancer region. In early Drosophila development, enhancers often have 10 – 20 TF binding sites for different transcription factors and different binding affinities, making accurate parameter estimation virtually impossible due to the large number of model parameters and potential parameter compensation.
To combat this issue, we have begun designing synthetic enhancers to allow us to mathematically model and fit parameter values on simple enhancers. This will allow us to decipher the basic underlying rules behind the interactions between TFs before testing the model on more complex endogenous enhancers involved in embryonic development.